Posterior over ontology nodes
The toy theorem panel shows how NuClass keeps probability mass on both internal and leaf nodes using a stop-at-node formulation.
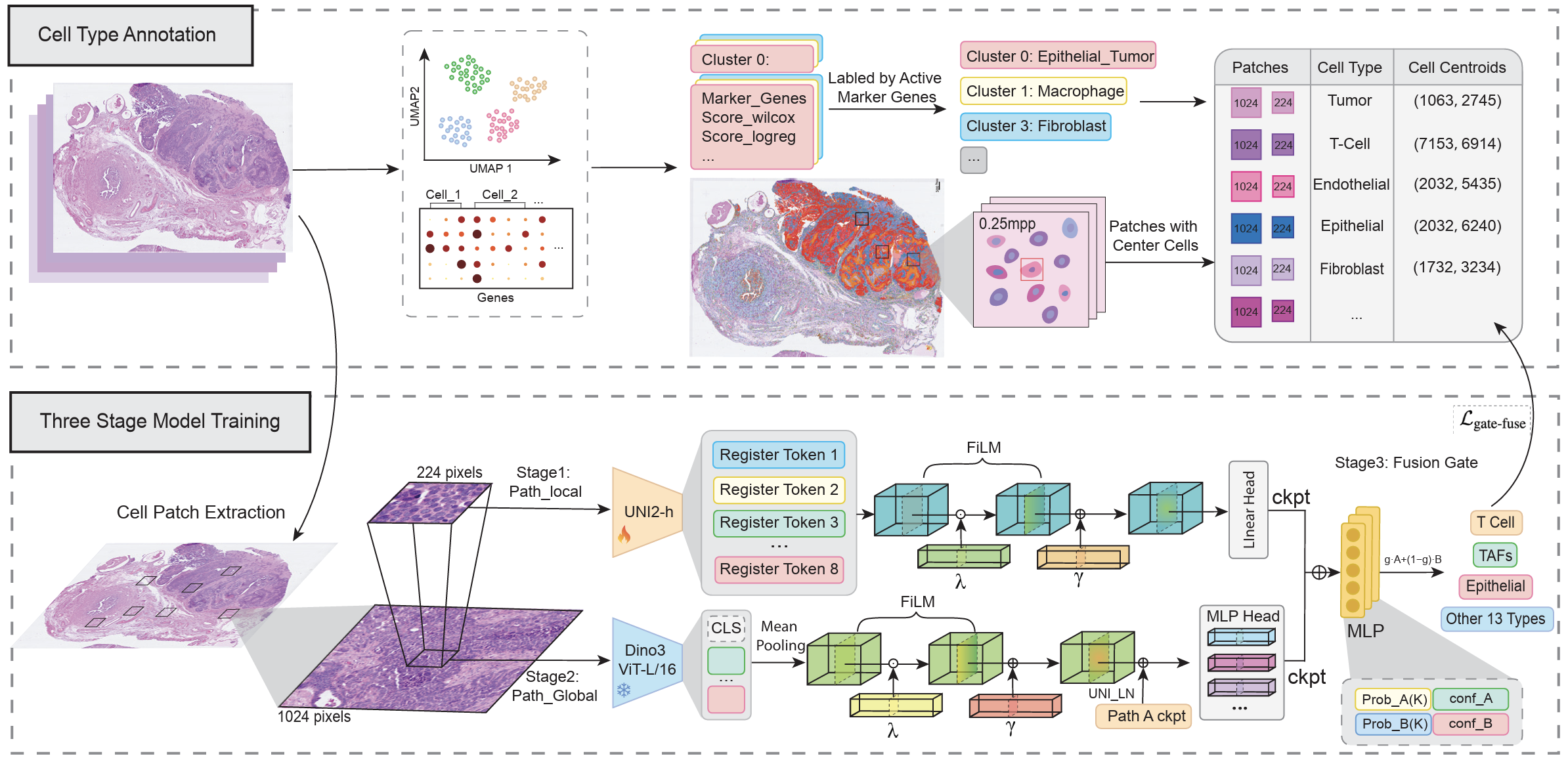
Adaptive multi-scale integration for robust cell annotation across histopathology and spatial imaging data.
This page combines the theorem intuition and the ontology explorer into one interface. The top section explains the training and Bayes-risk decision rule on a toy tree, and the second section lets you explore the larger NuClass cell hierarchy used for annotation.
The toy theorem panel shows how NuClass keeps probability mass on both internal and leaf nodes using a stop-at-node formulation.
Coarse labels become subtree events, so the likelihood aggregates all descendants that remain consistent with the observed label.
Prediction is based on Bayes risk under tree distance, which makes ontology-aware decisions different from plain posterior argmax.
The toy tree uses the ontology A -> {B, C}, C -> {D, E}. Adjust the logits to see how posterior mass changes, how a coarse label becomes a subtree likelihood during training, and how Bayes-risk decoding can choose a different prediction than posterior argmax.
This larger tree lets you inspect the cell hierarchy directly. You can change layouts, control visible depth, filter lineages, restrict to dataset nodes, and search for specific cell types while keeping the visual language aligned with the theorem panel above.
The proof PDF is embedded here for quick reference. If your browser blocks inline PDF rendering, use the preview or download buttons instead.
The same proof document is available for direct browser preview and local download from the buttons on the right.